Difference between revisions of "Tkt"
m (Reverted edits by 69.64.46.108 (talk) to last revision by 134.76.70.252) |
|||
| Line 41: | Line 41: | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
| − | + | <br/><br/> | |
| − | |||
| − | |||
| − | |||
| − | |||
= [[Categories]] containing this gene/protein = | = [[Categories]] containing this gene/protein = | ||
| Line 117: | Line 113: | ||
* '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=tkt_1919861_1921864_1 tkt] {{PubMed|22383849}} | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=tkt_1919861_1921864_1 tkt] {{PubMed|22383849}} | ||
| − | * '''Sigma factor:''' | + | * '''[[Sigma factor]]:''' |
* '''Regulation:''' | * '''Regulation:''' | ||
| Line 153: | Line 149: | ||
<pubmed>9924800 </pubmed> | <pubmed>9924800 </pubmed> | ||
==Original publications== | ==Original publications== | ||
| − | <pubmed>14651647,16493705, 19603213 9068642 16143417 15375635 19193632 </pubmed> | + | <pubmed>14651647,16493705, 19603213 9068642 16143417 15375635 19193632 23568212 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 11:57, 15 April 2013
- Description: transketolase
| Gene name | tkt |
| Synonyms | tktA |
| Essential | no |
| Product | transketolase |
| Function | pentose phosphate pathway |
| Gene expression levels in SubtiExpress: tkt | |
| Interactions involving this protein in SubtInteract: Tkt | |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 72 kDa, 4.803 |
| Gene length, protein length | 2001 bp, 667 aa |
| Immediate neighbours | ynzC, sirA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 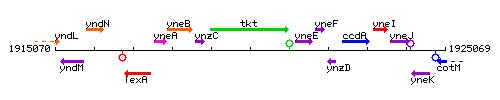
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed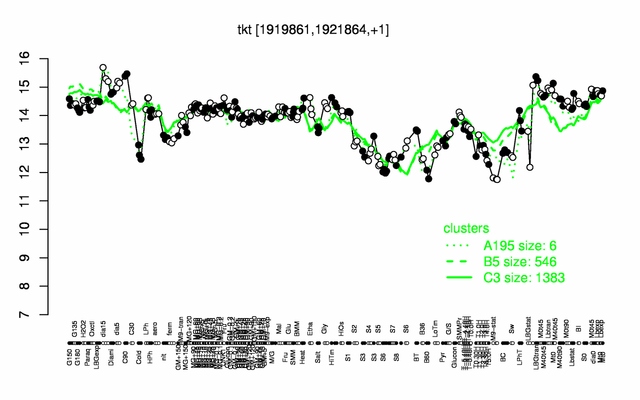
| |
Contents
Categories containing this gene/protein
carbon core metabolism, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU17890
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Sedoheptulose 7-phosphate + D-glyceraldehyde 3-phosphate = D-ribose 5-phosphate + D-xylulose 5-phosphate (according to Swiss-Prot)
- Protein family: transketolase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylated on ser/ thr/ tyr PubMed
- Cofactor(s): thiamine pyrophosphate
- Effectors of protein activity:
Database entries
- Structure: [3K95], complex with thiamine diphosphate, from B. anthracis
- UniProt: P45694
- KEGG entry: [3]
- E.C. number: 2.2.1.1
Additional information
Expression and regulation
- Operon: tkt PubMed
- Additional information:
Biological materials
- Mutant: BS4530 (aphA3), available in Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
G Schenk, R G Duggleby, P F Nixon
Properties and functions of the thiamin diphosphate dependent enzyme transketolase.
Int J Biochem Cell Biol: 1998, 30(12);1297-318
[PubMed:9924800]
[WorldCat.org]
[DOI]
(P p)
Original publications