Difference between revisions of "Lip"
(→References) |
(→References) |
||
| Line 133: | Line 133: | ||
[http://www.ncbi.nlm.nih.gov/pubmed/21815947 PubMed:21815947] | [http://www.ncbi.nlm.nih.gov/pubmed/21815947 PubMed:21815947] | ||
<pubmed> 15812018,12523966,18721749,12218047,16342303, 12951259 11583117 11491291</pubmed> | <pubmed> 15812018,12523966,18721749,12218047,16342303, 12951259 11583117 11491291</pubmed> | ||
| − | <pubmed> 18383241,19180538, 8396026 19883129 18840696 21477088 18053819 , </pubmed> | + | jetzzt 3 |
| + | <pubmed> 18383241,19180538, 8396026, </pubmed> | ||
| + | jetzt 4 | ||
| + | <pubmed> 19883129 18840696 21477088 18053819 , </pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 15:42, 20 November 2011
- Description: extracellular lipase
| Gene name | lip |
| Synonyms | lipA |
| Essential | no |
| Product | extracellular lipase |
| Function | lipid degradation |
| MW, pI | 22 kDa, 10.059 |
| Gene length, protein length | 636 bp, 212 aa |
| Immediate neighbours | ansZ, yczC |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 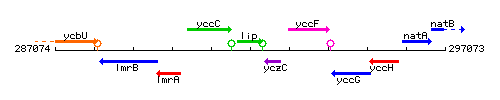
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU02700
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s): LipB
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure: 2QXT
- UniProt: P37957
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y. Mol Microbiol. 2011 81(6): 1459-1473. PubMed:21815947
jetzzt 3
jetzt 4
Zhong Ni, Peng Zhou, Xin Jin, Xian-Fu Lin
Integrating In Silico and In vitro approaches to dissect the stereoselectivity of Bacillus subtilis lipase A toward ketoprofen vinyl ester.
Chem Biol Drug Des: 2011, 78(2);301-8
[PubMed:21477088]
[WorldCat.org]
[DOI]
(I p)
Bo Chen, Zhen Cai, Wei Wu, Yunlong Huang, Juergen Pleiss, Zhanglin Lin
Morphing activity between structurally similar enzymes: from heme-free bromoperoxidase to lipase.
Biochemistry: 2009, 48(48);11496-504
[PubMed:19883129]
[WorldCat.org]
[DOI]
(I p)
Allison V Banse, Arnaud Chastanet, Lilah Rahn-Lee, Errett C Hobbs, Richard Losick
Parallel pathways of repression and antirepression governing the transition to stationary phase in Bacillus subtilis.
Proc Natl Acad Sci U S A: 2008, 105(40);15547-52
[PubMed:18840696]
[WorldCat.org]
[DOI]
(I p)
Eerappa Rajakumara, Priyamvada Acharya, Shoeb Ahmad, Rajan Sankaranaryanan, Nalam M Rao
Structural basis for the remarkable stability of Bacillus subtilis lipase (Lip A) at low pH.
Biochim Biophys Acta: 2008, 1784(2);302-11
[PubMed:18053819]
[WorldCat.org]
[DOI]
(P p)